The default test scripts run directly against the source code. dist : update the distribution js for npm etc.good for testing cytoscape in another project (with a "cytoscape": "file./path/to/cytoscape" reference in your project's package.json).watch:umd : automatically build prod umd bundle (no sourcemap, with babel).good for testing performance or for testing out of date browsers.watch:babel : automatically build lib for debugging (with sourcemap, with babel, a bit slower).served on or the first available port thereafter, with livereload on debug/index.html.good for general testing on debug/index.html.watch : automatically build lib for debugging (with sourcemap, no babel, very quick).docs : build the docs into documentation.build:esm : do the esm (ES 2015 modules) build. 
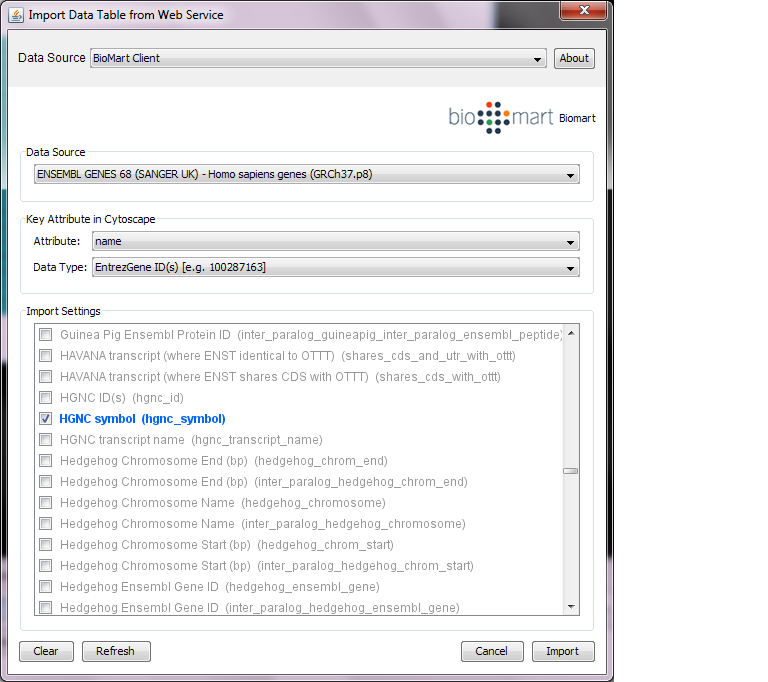
build:umd : do the umd (cjs/amd/globals) build.build:min : do the unminified build with bundled dependencies (for simple html pages, good for novices).build: do all builds of the library (umd, min, umd, esm).To cite Cytoscape.js in a paper, please cite the Oxford Bioinformatics issue:Ĭytoscape.js: a graph theory library for visualisation and analysisįranz M, Lopes CT, Huck G, Dong Y, Sumer O, Bader GDīioinformatics (2016) 32 (2): 309-311 first published online Septemdoi:10.1093/bioinformatics/btv557 (PDF) This allows for rapid releases of first- and third-party contributions. Get in touch with us by posting a GitHub discussion.įor the mechanics of contributing a pull request, refer to CONTRIBUTING.md.įeature releases are made monthly, while patch releases are made weekly. documentation, outreach), depending on your interests. features, testing) or non-technical roles (e.g. Would you like to become a Cytoscape.js contributor? You can contribute in technical roles (e.g. You can use the milestones to see what's currently planned for future releases. Roadmapįuture versions of Cytoscape.js are planned in the milestones of the Github issue tracker. You can find the documentation and downloads on the project website. More demos are available in the documentation. The Tokyo railway stations network can be visualised with Cytoscape:Ī live demo and source code are available for the Tokyo railway stations graph. Learn more about the features of Cytoscape.js by reading its documentation. If you only interested in interactions among given genes, you can do this by following “Loading a network” and you need to check the box of “Search Paths Between Genes”.Var cy = cytoscape ( ).It has three levels from 1 to 3 that mean users can choose different number of steps of expansion to find neighbors. There is interaction level drop-down list on the dialog box.Right click a node/edge->annotation->log in->change annotation set name->edit attributes from MiMI, add comments and URLs->save->log out.Select nodes and edges->right click node->SAGA->apply SAGA to chosen network->specify database and matching percentage->query.Click “sort” to sort the result with MEADįinding pathway matches for chosen network Right click an edge->get literature info from BioNLP.If you need to load a network, click here. Assuming that you already loaded a network.Getting literature information for selected edge (interaction) and sort the result Double clicking node/edge will direct you to MiMI webpage for that node/edge.Browse node/edge attributes with Cytoscape Node Attribute Browser and Edge Attribute Browser.Ī) Click Node Attribute Browser on Cytoscape panel (bottom right)->click icon ->check attributes name->choose nodes to see attributes.ī) Click Edge Attribute Browser to get edge information.Ī) Starting up Cytoscape->plugins->MiMI->MiMI Plugin->From File tab-> Drop down box ->specify species, molecule type etc Prepare a list of genes, see, for example mygene.txt.If you need to install Cytoscape, click here. Assuming that you already installed Cytoscape and MiMI Plugin.Finding pathway matches for chosen networkĬlick here to get instructions.Getting literature information for selected edge and sort the result.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
